Bioinformatics
Course code
MBR357 (undergrad), under the Department of Marine Biotechnology and Resources
Semesters taught
115-1 (offered every year in the first semester)
Course description
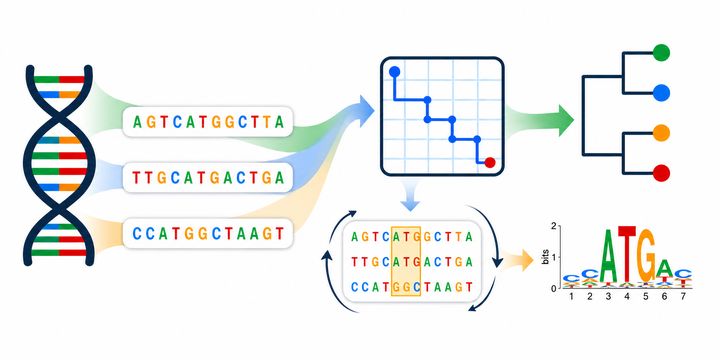
Bioinformatics integrates biology, statistics and computer science. This course is suitable for students who want to learn how to use computers to run analysis by building pipelines from existing software, or by designing a new algorithm. I will use some classical problems to demonstrate: sequence alignment, phylogeny reconstruction, motif finding, etc. The focus is not on the topics per se, but rather on how we formulate them as computationally solvable problems. The course is therefore designed to be general, with the hopes that the skills and concepts learned will be transferable across the vast number of topics in bioinformatics, and even outside of biology. Therefore, this course is also suitable for students with little biology background.
The first part of the course focuses on the practical data analysis. Students will learn how to work in a Linux virtual machine (using Github Codespaces):
- How to identify suitable software for a question
- How to install software for reproducible analysis
- How to format inputs and outputs so that different software tools can work together
- How to combine multiple software tools into a pipeline
- How to work with large files
These basics are best taught in a command line interface, inside a Linux environment so that students can better understand how a computer works.
The second part of the course focuses on learning the algorithms underlying these software. Students will learn:
- How many possible solutions does this problem have?
- How can we score a solution?
- How can we find the solution with the best score using the least computation?
- When the number of possibilities is too large, how can we find an approximate best solution?
Specifically, I will focus on the concepts (or their simplifications) of Dynamic Programming (DP), Maximum Likelihood Estimation (MLE), and Expectation Maximization (EM), etc. I will spend a significant amount of time on DP, building from the basic concepts of recursion and graph representation.
In the last part of the course, I will briefly introduce students to the concept of deep learning (DL), as it has become increasingly important in bioinformatics. The goal here is to connect the optimization methods students learned in the previous part to how DL models works with gradient descent.
Prerequisite
- Biological knowledge: Students only need to know that DNA is the genetic material of living organisms and that it consists of four letters: A, G, T, and C. That is all.
- Programming experience: Prior programming experience is NOT required, but it will be helpful. In the first part of the course, I will teach the necessary computational skills from scratch. In the second part, the focus will shift toward problem-solving by hand. Coding is less central in the age of AI; problem formulation and understanding algorithms are more important.
Syllabus
- to be updated